Enzymes for the biodetoxification of DON

The safe and efficient removal of the mycotoxin deoxynivalenol is a pressing issue in terms of ensuring food safety. Enzymes, isolated from microorganisms with DON detoxification activity, offer promising alternatives to alleviate this problem. However, some key challenges must be addressed before employing these enzymes for industrial applications.
Deoxynivalenol (DON) is a toxic secondary metabolite produced by filamentous fungi of the genus Fusarium. DON contamination continues to plague agricultural commodities globally. In fact, in a 10-year survey of global mycotoxin contaminations in grain samples, DON appeared to be the leading mycotoxin contaminant alongside other Fusarium mycotoxins such as fumonisins and zearalenone. Due to climate change, abiotic stresses such as CO2, temperature and humidity may provide a conducive environment for Fusarium growth, thus further exacerbating contamination of food by DON.
Regulating and mitigating DON levels is key to ensuring food safety and food security. Toxicology studies indicate that acute exposure to DON can result in emesis, while in the long term, it can lead to gastrointestinal ailments, compromise the immune system and cause negative reproductive effects.
Despite best efforts to monitor and control DON levels at the farm level, post-harvest measures such as physical and chemical decontamination are inevitably required. Unfortunately, these measures are only marginally effective, may introduce new toxic compounds, and compromise the nutritional quality and palatability of the finished product.
Learning from microbes: “The best defence is a good offence”
Biodetoxification, which is a growing field in mycotoxin research, employs novel microbes or their enzymes to convert DON to a less toxic form. Microbes have evolved an arsenal of novel defences to counteract the inherent toxicity of mycotoxins such as DON.
At the molecular level, DON binds tightly to ribosomes, inhibiting protein synthesis by these cellular molecules. The key structural features contributing to these interactions are the C3 hydroxyl group and the C12-13 epoxide ring of DON . Microbes use several strategies to modify these functional groups including de-epoxidation, oxidation, epimerization, hydrolysis, acetylation, hydroxylation, and glycosylation. Although these microbes can be used directly to detoxify DON, isolating the responsible enzymes offers the advantage of specificity, thus preventing unwanted side reactions catalysed by other microbial enzymes.
One promising pathway, known as the DON epimerisation (Dep) system was first extensively studied in the bacterium Devosia mutans 17-2-E-8, which detoxifies DON by converting it to the isomer, 3-epi-DON (Figure 1) . This impedes binding to the ribosome and toxicology assays indicate that 3-epi-DON is at least 50 times less toxic than DON. Livestock such as pigs, which have a low tolerance to DON in their diet, did not show any adverse effects when fed diets supplemented with 3-epi-DON .
Figure 1 – The Dep pathway in Devosia mutans 17-E-2-8.
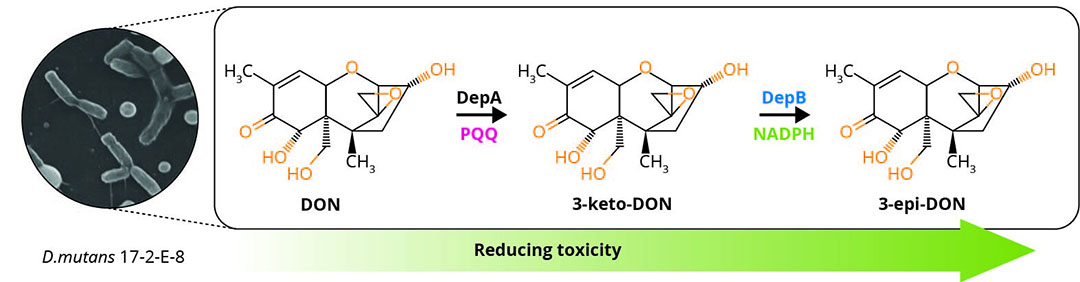
Key players of the Dep system: DepA and DepB
Epimerisation involves the activity of 2 enzymes.
DepA: oxidides DON at the C3 hydroxyl position to produce the compound 3-keto-DON.
DepB: then takes this compound, finally transforming it into 3-epi-DON. DepA possesses a high catalytic efficiency with DON at around 13,000 M-1 s-1 while DepB’s catalytic efficiency is 4670 M-1 s-1 with 3-keto-DON.
The Dep system is versatile and could potentially be used for the detoxification of other mycotoxins although more research on this is required. A recent study highlighted for the first time the fact that DepB can detoxify the mycotoxin patulin to a less toxic product: ascladiol. However, its catalytic efficiency with patulin is quite low; only 15 M-1s-1, which leaves significant room for improvement. Dep enzymes are also active in relation to the masked mycotoxin, including 15-acetyl DON.
Applications of Dep enzymes
DON occurrence in maize by-products can pose a serious hazard for the livestock industry. Maize gluten feed, maize steep liquor and dried distillers grains with solubles (DDGS) are nutritionally rich and often used to supplement livestock diets. Unfortunately, DON levels in these feed sources can exceed prescribed thresholds. In cases such as these, applying Dep enzymes as processing aids could reduce DON levels. Alternatively, the enzymes could be used to detoxify DON in animal feed to reduce the bioavailability of the mycotoxin when ingested by livestock.
Challenges
A caveat of the Dep system is the fact that both enzymes require cofactors. These are additional non-proteinaceous compounds that are necessary for the enzyme to carry out its function. For the Dep system to work, DepA requires pyrroloquinoline quinone or PQQ, while DepB requires NADPH.
Figure 2 – X-ray structure of DepB with NADPH showing a close up of potential residues involved in NADPH specificity.
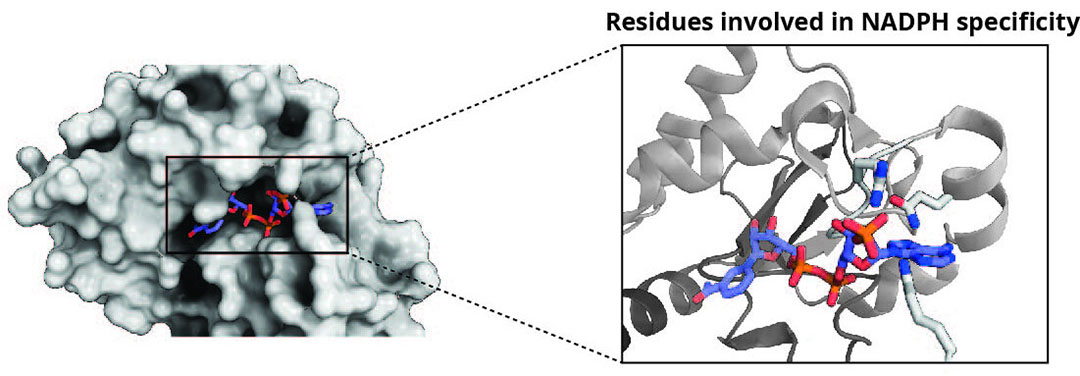
Biochemical characterisation of DepB has revealed that it could also use NADH to convert 3-keto-DON to 3-epi-DON, although its catalytic efficiency is 40 times lower than when NADPH was used. However, the advantages are that NADH is more stable and much cheaper than NADPH. By using protein engineering approaches, it may be possible to alter the enzymes cofactor preference for NADH. Analysis of the x-ray crystal structure of DepB was key to identifying residues potentially involved in NADPH specificity (Figure 2). Future steps will involve site-directed mutagenesis at these locations to improve the enzymes utilisation of NADH and keep costs at a minimum. Moreover, NADH can also be recycled with the addition of another enzyme, formate dehydrogenase (FDH), which only requires the cheap substrate formate and produces the by-product COzsa. This technique has been demonstrated successfully for the enzymatic generation of the sweetener, allitol.
A second issue that needs to be addressed is the pH sensitivity of the Dep system. The pH of animal feed products can range between 3.6 to 5 which may affect the activity of the enzymes. Currently the Dep system works optimally at a neutral pH, hence the pH of animal feed may need to be raised prior to using these enzymes for in situ detoxification of DON.
In conclusion – exciting new opportunity
In conclusion, the Dep system provides an exciting new opportunity to level the playing field with DON. However, the technology could still benefit from protein engineering to reduce costs associated with expensive cofactors. Alternatively, if using pure enzymes by themselves is too expensive, the enzymes could be co-expressed in other host microorganisms rather than the original microbial strain. This has several advantages such as a controlled microenvironment for enzymes to function, no requirements for additional cofactors and moreover this approach is cheaper and less labour-intensive than producing purified enzymes.
“The financial support from Agriculture and Agri-Food Canada and Grain Farmers of Ontario through the Industry-led Research and Development Project (J-002433) is acknowledged and much appreciated.”
References upon request.











